Research Articles
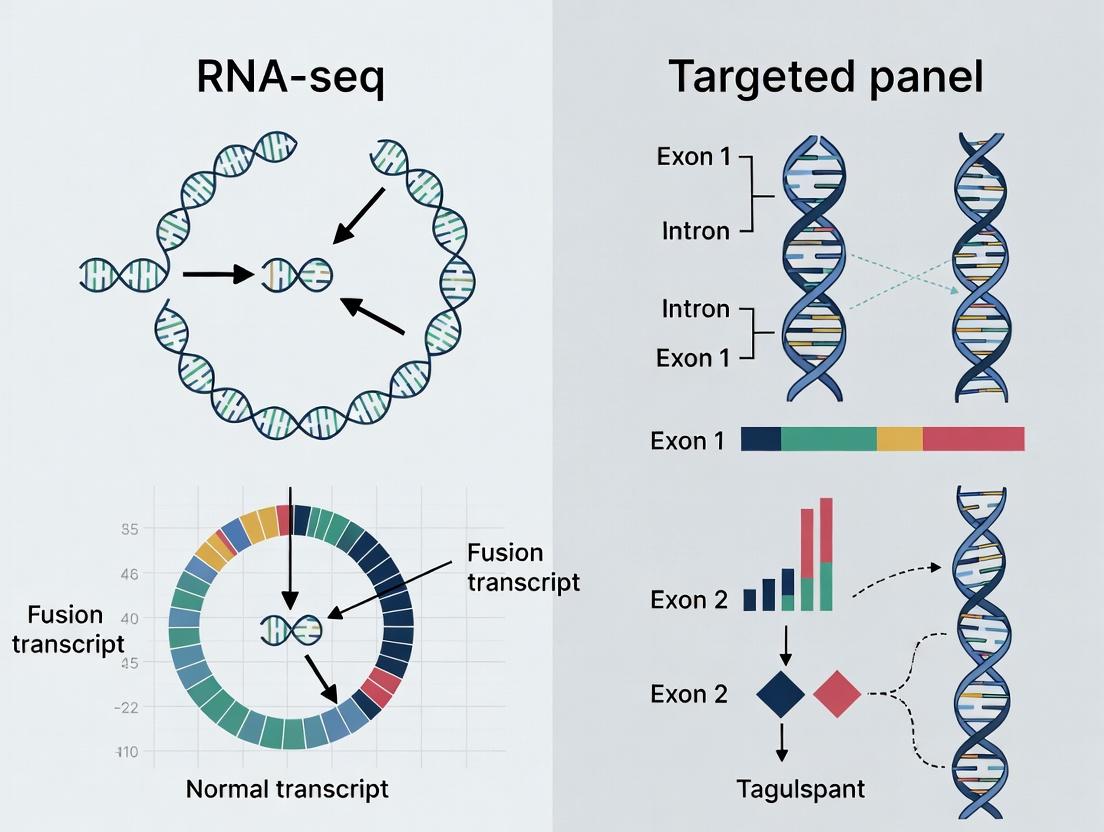
Fusion Detection in Cancer: Choosing Between RNA-seq and Targeted Panels for Precision Oncology
This article provides a comprehensive comparative analysis of RNA sequencing (RNA-seq) and targeted gene fusion panels for oncogenic fusion detection in cancer research and drug development.
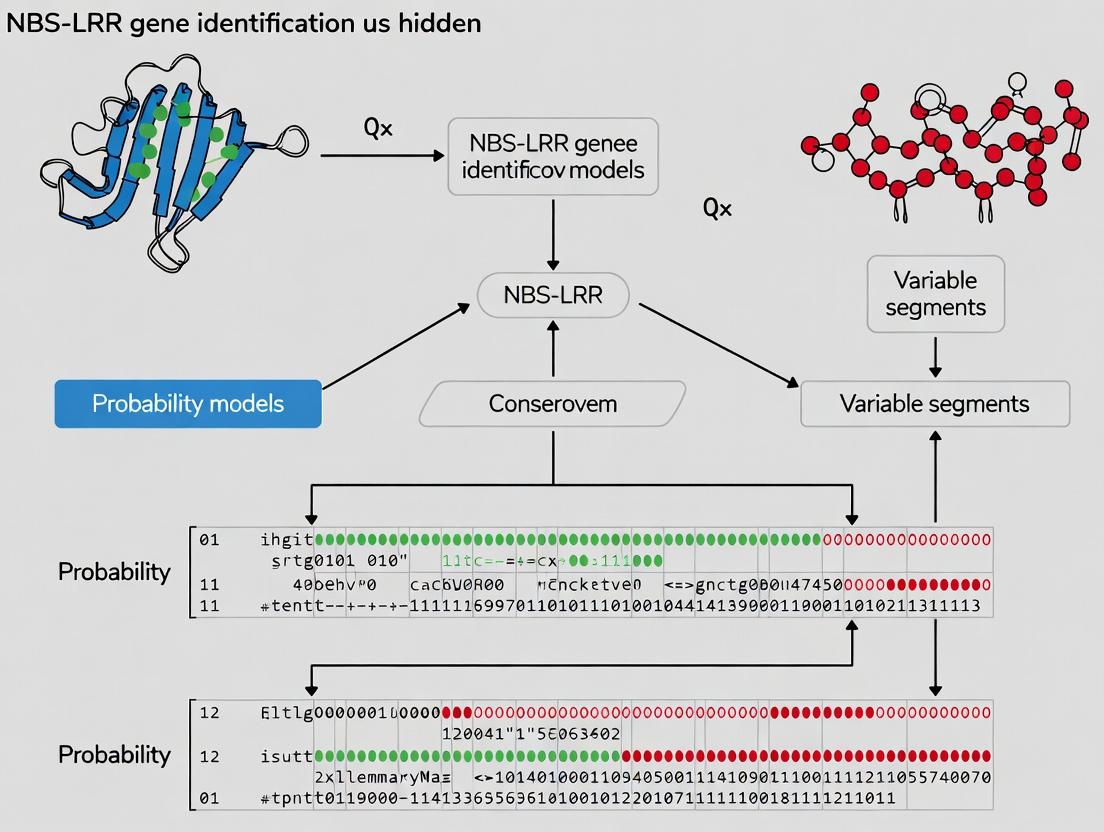
Unlocking Plant Immunity Genes: A Comprehensive Guide to NBS-LRR Gene Identification with Hidden Markov Models
This article provides a complete methodology for the identification of Nucleotide-Binding Site Leucine-Rich Repeat (NBS-LRR) genes, the largest class of plant disease resistance (R) genes.
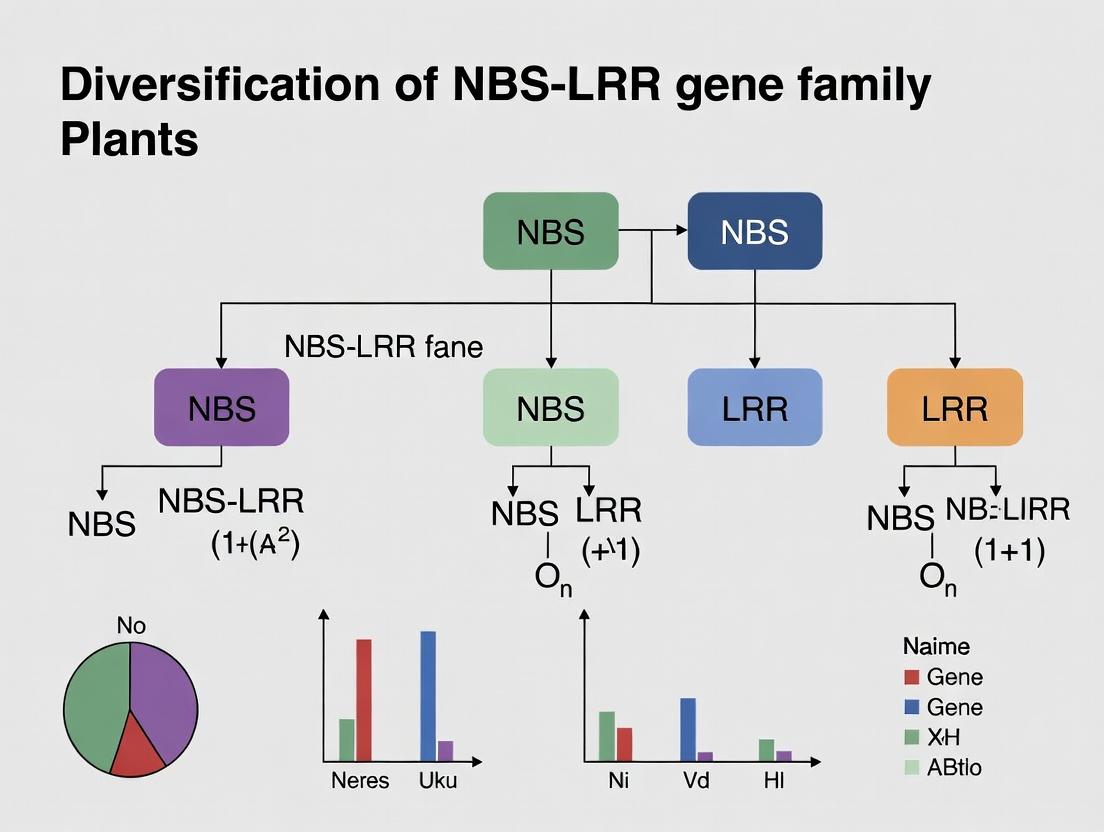
NBS-LRR Genes in Plants: Evolution, Mechanisms, and Biomedical Applications
This comprehensive review explores the diversification of the NBS-LRR gene family, the cornerstone of plant innate immunity.
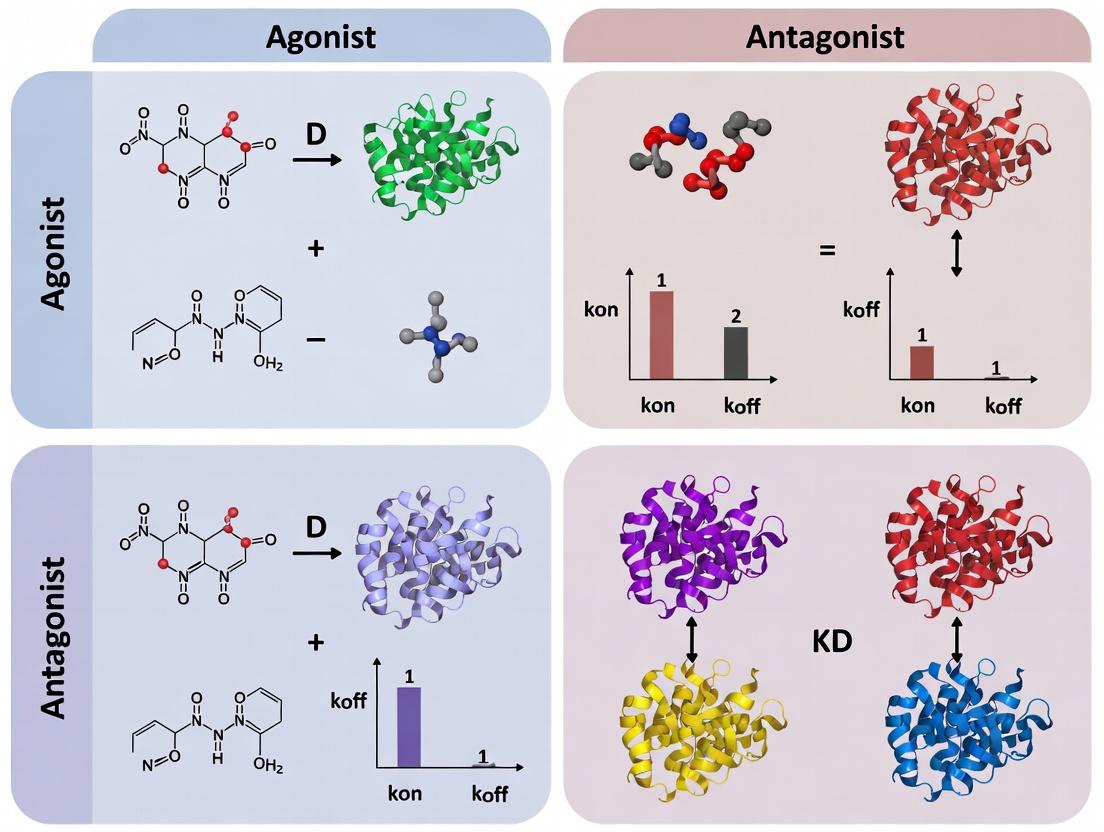
Decoding Drug Action: A Comparative Analysis of NBS-LRR Binding Kinetics for Agonists vs. Antagonists
This article provides a comprehensive guide for researchers and drug development professionals on the critical comparison of nucleotide-binding site-leucine-rich repeat (NBS-LRR) protein binding kinetics between agonists and antagonists.
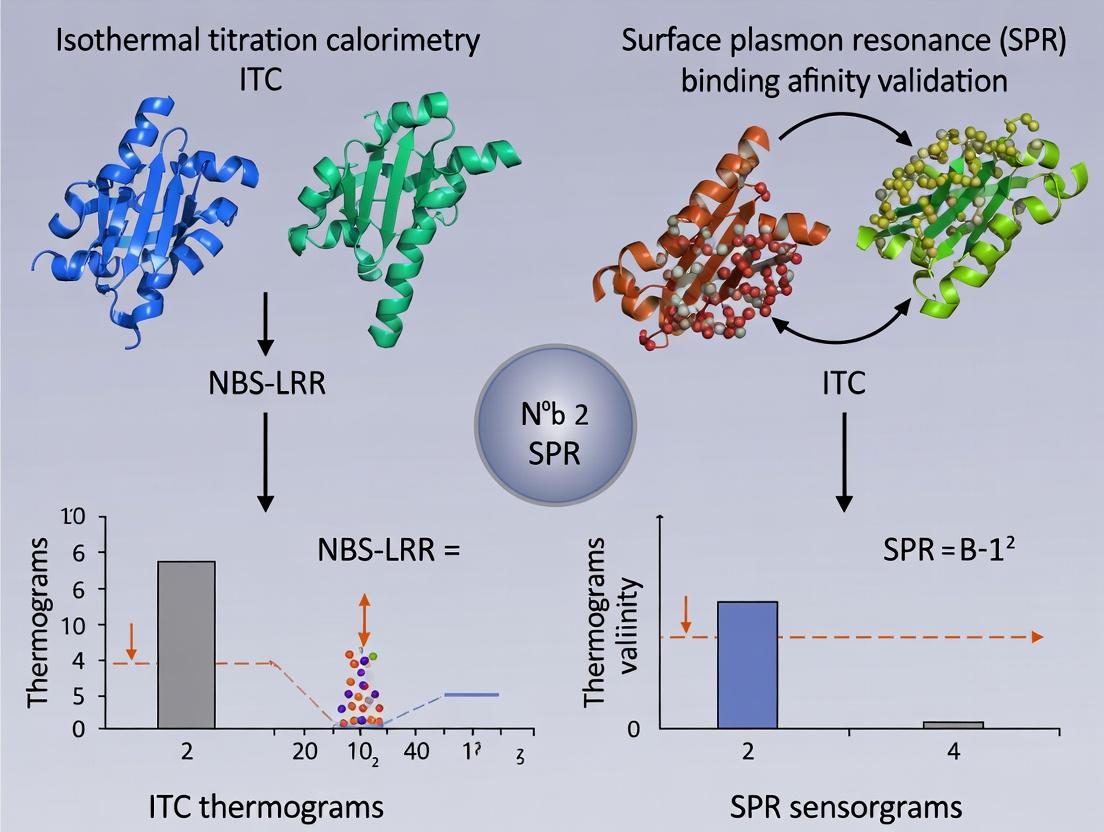
ITC vs SPR for NBS-LRR Binding Affinity: A Comparative Guide for Biophysical Validation
This article provides a comprehensive comparison of Isothermal Titration Calorimetry (ITC) and Surface Plasmon Resonance (SPR) for validating the binding affinity of NBS-LRR immune receptor proteins with their pathogen-derived ligands.
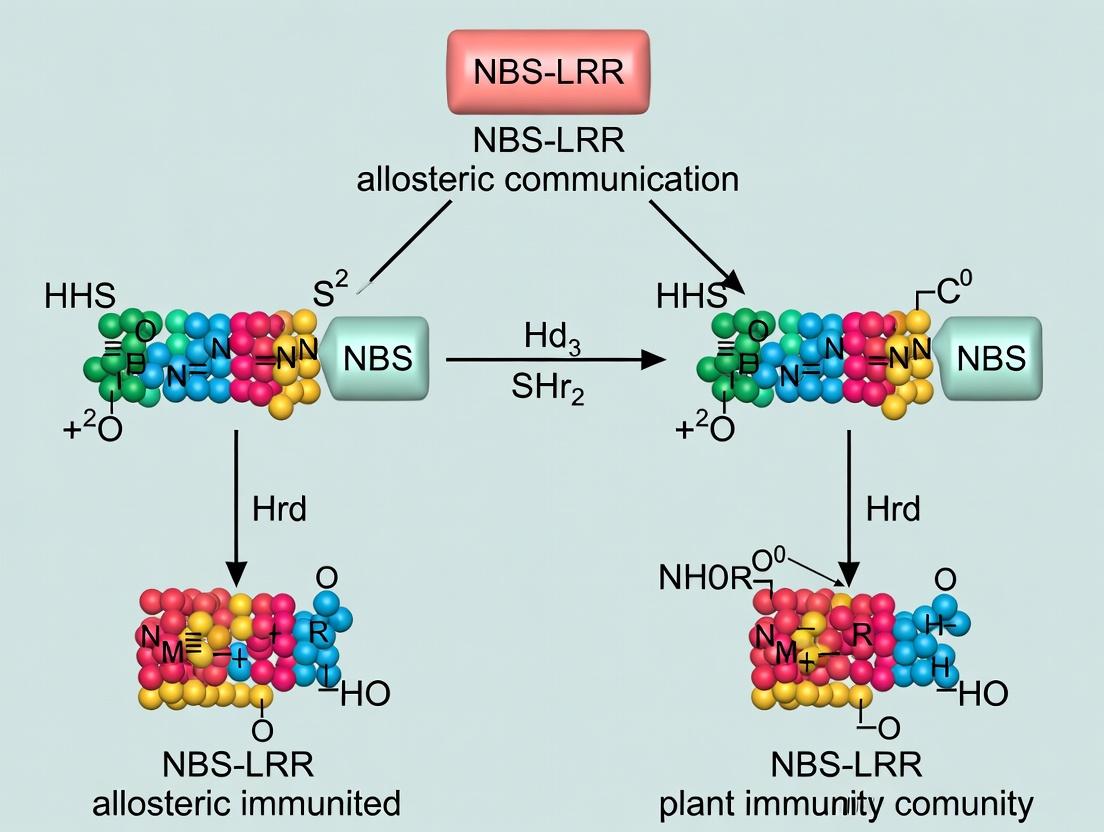
Unlocking the Molecular Switch: Allosteric Communication in NBS-LRR Immune Receptors for Drug Discovery
This comprehensive review synthesizes current research on the structural dynamics and allosteric communication networks within NBS-LRR (Nucleotide-Binding Site-Leucine-Rich Repeat) immune receptors.
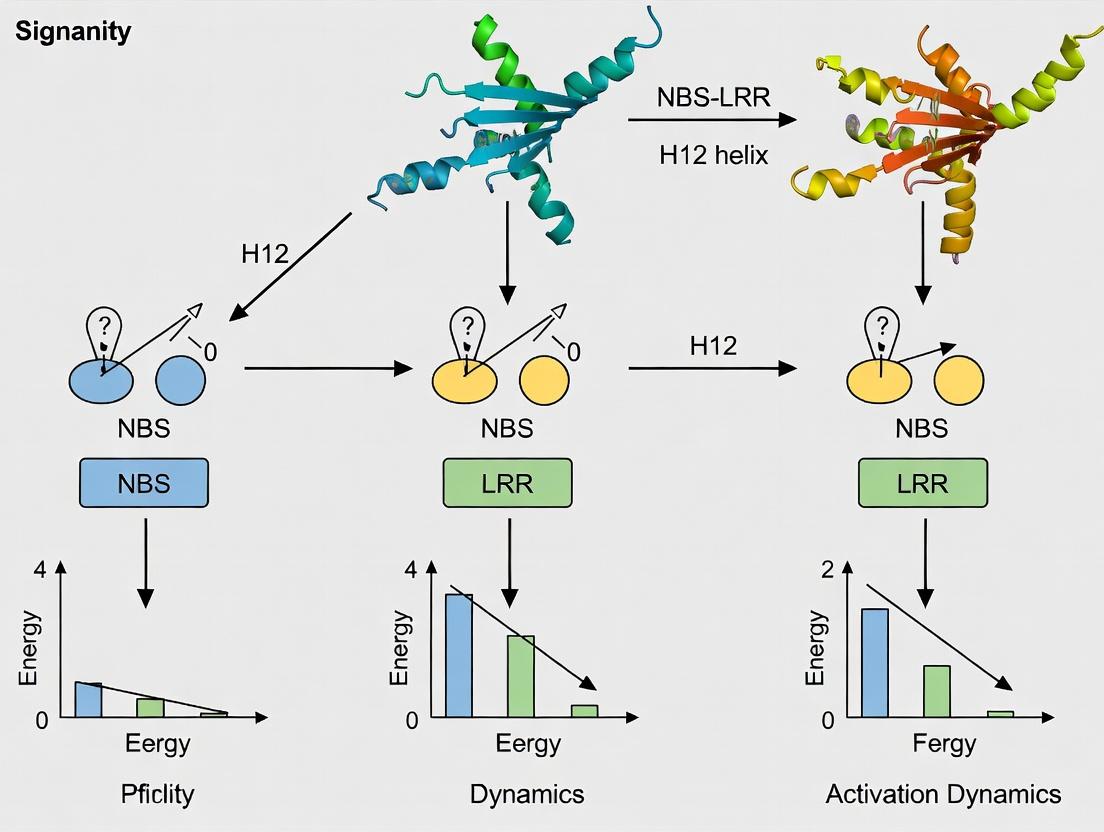
Unlocking Immune Activation: The Critical Role of H12 Helix Dynamics in NBS-LRR Proteins
This article provides a comprehensive analysis of the activation mechanisms of Nucleotide-Binding Site Leucine-Rich Repeat (NBS-LRR) proteins, focusing on the pivotal conformational dynamics of the H12 (Helix 12) motif.
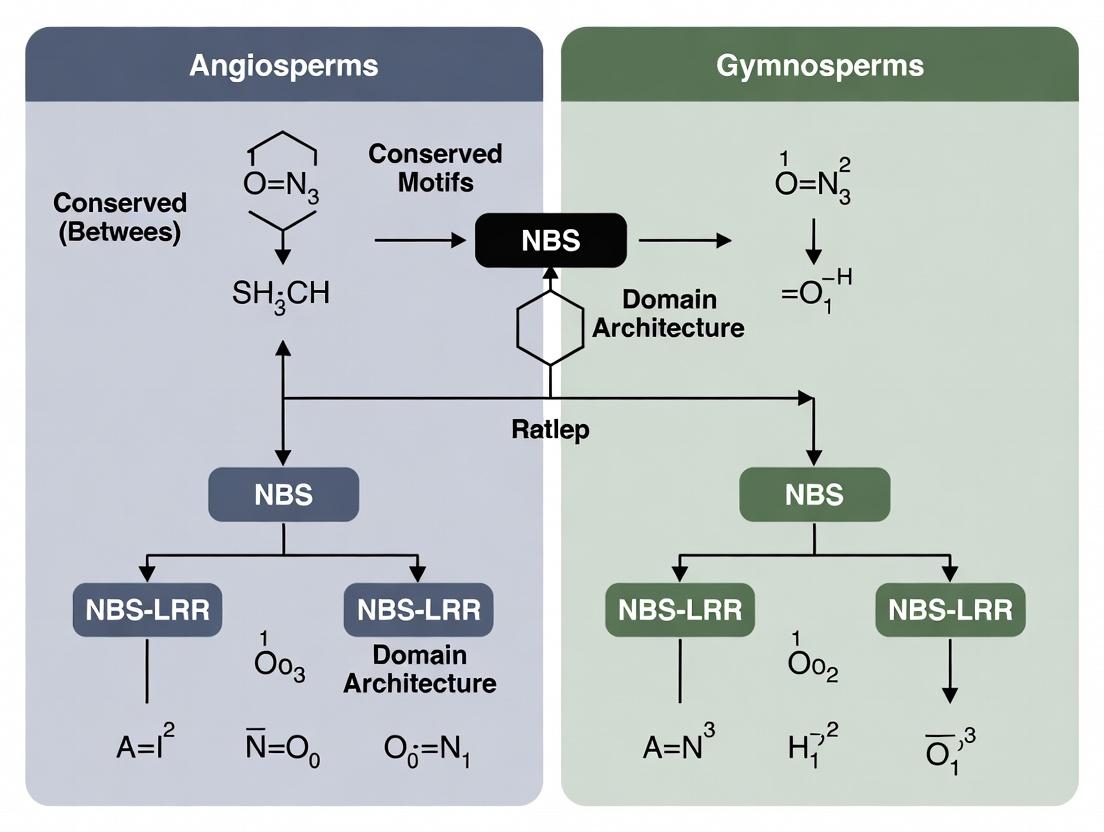
NBS Gene Family Evolution: Decoding Divergence Patterns Between Angiosperms and Gymnosperms for Biomedical Insights
This article provides a comprehensive analysis of the evolution of the Nucleotide-Binding Site (NBS) gene family, a crucial component of plant innate immunity, across angiosperms and gymnosperms.
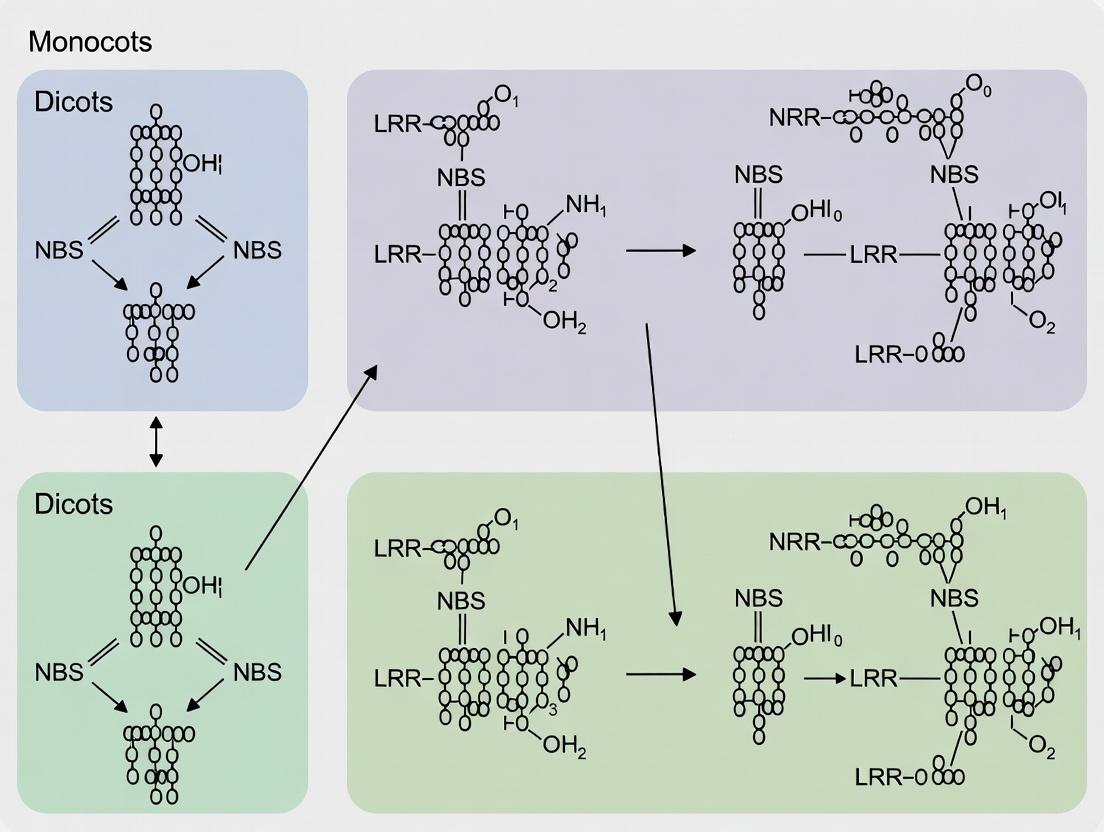
NBS Gene Family Evolution: A Comparative Genomics Analysis of Resistance Genes in Monocots vs. Dicots
This article provides a comprehensive analysis of the nucleotide-binding site leucine-rich repeat (NBS-LRR) gene family, the primary class of plant disease resistance (R) genes, through a comparative genomics lens.
NBS Gene Dynamics: Expansion, Contraction, and Adaptive Evolution in Plant Genomes
This article provides a comprehensive analysis of Nucleotide-Binding Site (NBS) gene family dynamics in plant genomes, focusing on the evolutionary forces driving their expansion and contraction.